Chromatin Immunoprecipitation Sequencing (ChIP-seq)
Introduction to ChIP-seq
Chromatin Immunoprecipitation Sequencing universally known as ChIP-seq is one of the most influential techniques in modern genomics.
Chromatin Immunoprecipitation Sequencing (ChIP-seq) is a genome-wide method used to identify protein-DNA binding events, histone modifications, and chromatin-associated regulatory mechanisms with high accuracy.
Combining immunoprecipitation with next-generation sequencing (NGS), ChIP-seq offers a precise, scalable approach for mapping epigenomic landscapes.
What Makes ChIP-seq Important ?
ChIP-seq enables researchers to uncover where specific proteins bind on DNA, including:
- Transcription factors
- Histone modifications
- Chromatin-associated proteins
- Epigenetic regulators
Genome-wide protein occupancy patterns
By revealing these interaction sites, ChIP-seq helps scientists understand gene regulation, chromatin architecture, and cellular identity on a genomic scale. NCBI
The ChIP-seq Workflow
ChIP-seq relies on a multi-stage experimental design integrating molecular preparation, immunoprecipitation, sequencing library generation, and computational analysis.
1-Crosslinking of Protein-DNA Complexes
Crosslinking stabilizes protein–DNA interactions in vivo.
Formaldehyde is commonly used due to its reversible nature and short crosslink distances. Advanced protocols may use :
- Formaldehyde alone (most common).
- Dual crosslinking (e.g., DSG + formaldehyde).
- UV-crosslinking for nucleic acid–protein complexes.
2-Chromatin Extraction and Fragmentation
Once crosslinked, cells undergo lysis and chromatin extraction.
Two primary fragmentation strategies exist :
Mechanical Fragmentation
- Sonication using focused ultrasonication instruments
- Generates fragments between 100 – 600 bp
- Ideal for transcription factor ChIP-seq
Enzymatic Digestion
- Micrococcal nuclease (MNase)
- Produces nucleosome-sized fragments
- Preferred for histone modification studies
3-Immunoprecipitation Using Highly Specific Antibodies
The immunoprecipitation step enriches DNA fragments bound to the protein of interest.
Key variables include :
- Antibody specificity
- Epitope affinity
- Batch consistency
- Chromatin input quality
Magnetic beads coated with Protein A/G are commonly used.
4-Reverse Crosslinking and DNA Purification
After immunoprecipitation, the following steps occur :
- Reverse crosslinking (typically at high temperature)
- Protease treatment
- Purification of ChIP-DNA
This yields high-enrichment, low-background DNA suitable for sequencing.
Library Preparation for Next-Generation Sequencing
Library construction involves :
- End repair
- A-tail addition
- Adapter ligation
- PCR amplification
- Size selection
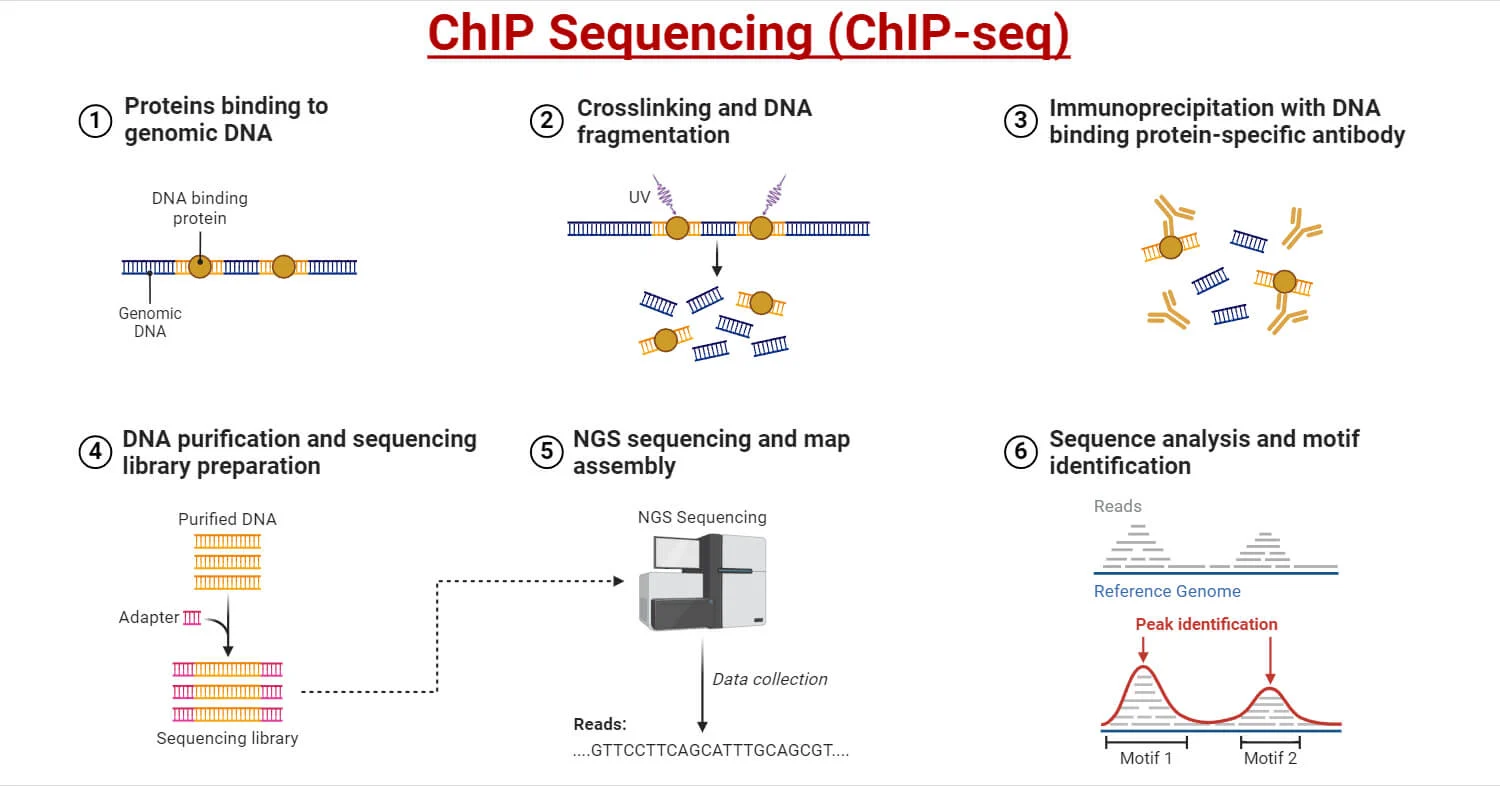
Bioinformatics Analysis
Read Quality Control (QC)
Standard QC involves :
-Phred score evaluation
-Adapter trimming
-Removal of low-quality reads
Alignment to Reference Genome
Reads are mapped to reference genomes using high-performance aligners :
-BWA-MEM
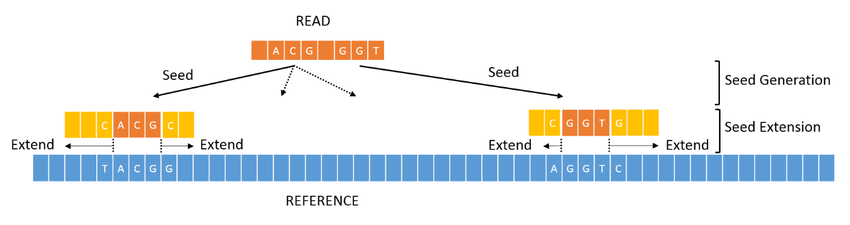
-Bowtie2

-HISAT2

Peak Calling : Identifying Protein Binding or Histone Mark Enrichment
Peak calling is a cornerstone of ChIP-seq analysis.
Tools include :
-MACS2 (Model-based Analysis of ChIP-seq). NCBI

-SICER (useful for broad histone marks)
-HOMER
Normalization and Control Correction
Input DNA or IgG controls allow removal of sequencing bias.
Normalization strategies include :
-RPM (Reads Per Million)
-RPKM
-Spike-in normalization
-Background subtraction
Motif Discovery and Functional Annotation
Once peaks are identified, downstream analysis includes :
-Motif enrichment analysis
-Gene ontology (GO) enrichment
-Pathway association analysis
-Peak annotation to promoters, enhancers, or intergenic regions
These steps bridge ChIP-seq data with functional genomic interpretation.
Applications of ChIP-seq
Transcription Factor Binding Maps
Transcription factors modulate gene expression.
ChIP-seq identifies :
- motif preferences
- co-factor interactions
- regulatory networks
- genomic occupancy patterns.
Histone Modification Profiling
Different histone marks correspond to specific regulatory states :
-H3K4me1 : enhancers

-H3K4me3 : active promoters

-H3K27ac : active regulatory regions

-H3K27me3 : repressed chromatin

-H3K9me3 : heterochromatin
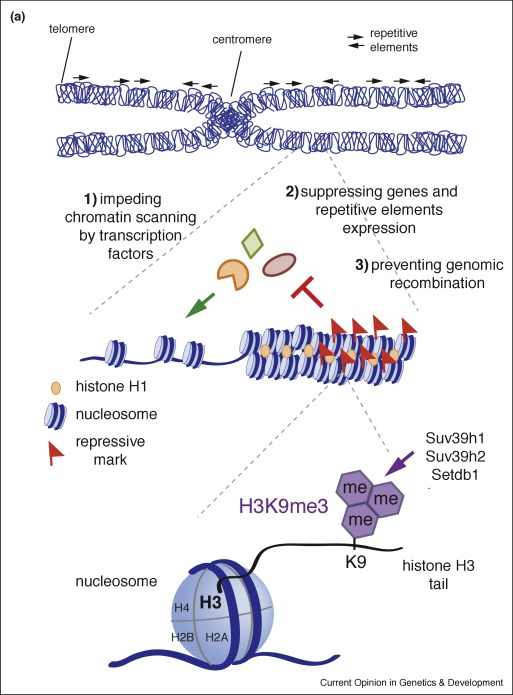
Chromatin State Dynamics
When combined with :
-ATAC-seq

-RNA-seq

-Hi-C

-DNA methylation assays

Comparative Genomics
Comparisons across species, conditions, or development stages provide insights into :
- evolutionary conservation
- epigenomic divergence
- environmental adaptation
Strengths and Limitations of ChIP-seq
Strengths
- High resolution
- Genome-wide scope
- Compatibility with many proteins
- Strongly supported by bioinformatics tools
- Generates interpretable peak landscapes
Limitations
- Dependent on antibody quality
- Requires significant sequencing depth
- Computationally intensive
- May generate false positives without proper controls

